Part 3: Energetics model
The specification of an energetics model is very important in detailed KMC simulations, as it captures the adsorbate-adsorbate lateral interactions responsible for ordering in the adsorbate overlayer. Lateral interactions also affect the activation barriers of elementary events, by stabilising or destabilising the initial, transition, or final state of the event.
The ZGB model does not include lateral interactions between adsorbates and the rates of events are explicitly defined (so activation energies will be taken as 0). We will, however, discuss, for the sake of completeness how to set up a simple model for the energetics, including only the binding energies of individual adsorbates (no interactions). We will assume a binding energy of 1.3 eV for CO and 2.3 for O.
Zacros implements a general cluster-expansion Hamiltonian approach for the energetics. In this approach, the energy of a given configuration is computed by summing up single-body contributions, as well as two-body, three-body etc. interaction terms. Thus, we have to define the individual patterns contributing to the overall energy, and the individual energy contribution of each pattern. This is done in the energetics_input.dat file which starts with keyword energetics, followed by one or more blocks of instructions contained within keywords cluster … end_cluster. The parsing of the file finished when Zacros encounters the keyword end_energetics.
To define correctly the patterns in Zacros, it is often helpful to make a schematic of the binding configuration wherein you clearly number the sites, adsorbate molecules and dentates thereof. This drawing process can be accomplished by following these steps:
- Draw all the sites involved in the pattern.
- If more than one sites are involved, draw all the links between the sites indicating which sites are neighbours.
- Number all the sites in the pattern, as s1, s2,…
- Draw all the adsorbate entities participating on the pattern, clearly indicating the species type for each entity.
- Draw the bonds between adsorbates and sites, to clearly indicate which adsorbate occupies which site (or sites, if there are multidentate adsorbates).
- Number all adsorbates (entities) as e1, e2,…
- Number all dentates as d1,1, d1,2, …, d2,1, d2,2,… The first index in this numbering system is the entity number, and the second is the dentate number.
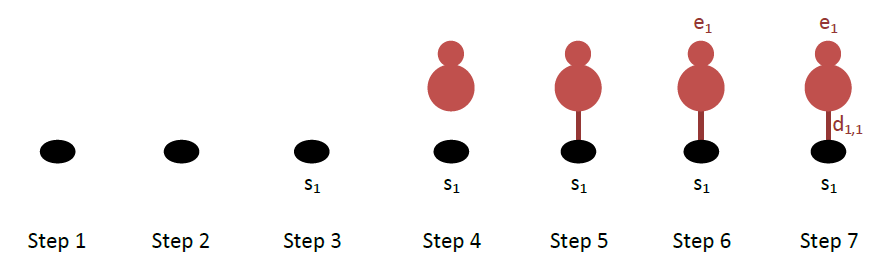
Let us do this for CO: this molecule binds to a single site. The drawings per step are shown below:
The Zacros input for the CO binding pattern starts with the keyword cluster followed by the name of the pattern:
cluster CO_point
Then, the number of sites involved is defined as follows:
sites 1
In this case we have only one site, so there are no links between sites. If we had 2 or more sites, we would have to define the neighbouring structure of the pattern using the keyword neighboring, which however in our case is omitted.
The most important part of the cluster definition is the description of the occupancies of the sites, which is done with the lattice_state keyword. This keyword is followed by as many lines of input as the number of sites in the pattern (in this case only one). Each line contains three words: the first is the number of the entity/adsorbate occupying the corresponding site, the second is the species name occupying that site, and the last word is the dentate number with which the entity binds to that site. In our case (see above figure), site number 1 is occupied by dentate 1 of entity number 1, which is a CO*. Therefore:
lattice_state
1 CO* 1
The last two things we need to define here are the graph multiplicity and the energy contribution of the cluster (also referred to as effective cluster interaction). The multiplicity is defined by the optional keyword graph_multiplicity, whereas the energy contribution is specified with the keyword cluster_eng. If graph_multiplicity is absent, a default value of 1 is used for the multiplicity. The latter exists for cross-compatibility with other KMC software codes: when Zacros detects this pattern on the lattice, it will add a contribution of cluster equal to the cluster energy divided by the multiplicity. This is useful, for instance, in the case of a symmetric pairwise pattern, which will be detected twice by the Zacros algorithm. A multiplicity of 2 will correct for this double-counting. In our case, we have a single site pattern so things are rather simple:
graph_multiplicity 1
cluster_eng -1.3
The cluster specification block ends with the keyword end_cluster.
Similarly, the corresponding block for the adsorbed O cluster is:
cluster O_point
sites 1
lattice_state
1 O* 1
graph_multiplicity 1
cluster_eng -2.3
end_cluster